
・Basic Research・
Calpastatin
participates in the regulation of cell migration in BAP1-deficient uveal
melanoma cells
Han Yue1,2, Feng-Xi
Meng1,2, Jiang Qian1,2, Bin-Bin Xu1,2, Gang
Li2,3, Ji-Hong Wu2,3
1Department
of Ophthalmology, Eye & ENT Hospital of Fudan University, Shanghai 200031,
China
2Shanghai Key
Laboratory of Visual Impairment and Restoration, Fudan University, Shanghai
200433, China
3Experimental
Research Center, Eye & ENT Hospital of Fudan University, Shanghai 200031,
China
Co-first
authors: Han Yue and Feng-Xi Meng
Correspondence
to: Jiang Qian. Department of Ophthalmology, Eye & ENT
Hospital of Fudan University, 83 Fenyang Rd, Shanghai 200031, China.
qianjiang58@hotmail.com
Received:
Abstract
AIM: To detect how BRCA-associated protein 1 (BAP1) regulates cell migration
in uveal melanoma (UM) cells.
METHODS: Wound healing and transwell assays were performed
to detect UM cell migration abilities. Protein chip, immunoprecipitations and
surface plasmon resonance analyses were applied to identify BAP1 protein
partners. Western blot and calpain activity assays were used to test the
expression and function of calpastatin (CAST).
RESULTS: CAST protein was confirmed as a new BAP1 protein
partner, and loss of BAP1 reduced the expression and function of CAST in UM
cells. The overexpression of CAST rescued the cell migration phenotype caused
by BAP1 loss.
CONCLUSION: BAP1 interacts with CAST in UM cells, and CAST and
its subsequent calpain pathway may mediate BAP1-related cell migration
regulation.
KEYWORDS: uveal
melanoma; BRCA-associated protein 1; calpastatin; cell migration
DOI:10.18240/ijo.2019.11.03
Citation: Yue H, Meng FX, Qian J, Xu BB, Li
G, Wu JH. Calpastatin participates in the
regulation of cell migration in BAP1-deficient uveal melanoma cells. Int J
Ophthalmol 2019;12(11):1680-1687
INTRODUCTION
Uveal
melanoma (UM) is the most common intraocular primary malignancy in adults. The
estimated incidence of this disease is 5 or 6 cases per million per year[1-2], and almost half of the patients
will die from metastases within approximately 10y[3-4]. The most predominant locations for metastases are the
liver (89%), followed by the lungs (29%) and bone (17%)[5].
Thus, preventing metastasis at an early stage and discovering the underlying
mechanism of micrometastasis are important topics.
In 2010,
Harbour et al[6] reported that bap1,
the gene encoding BRCA-associated protein 1 (BAP1), was mutated in
approximately 84% of metastatic UMs and indicated that germline bap1
mutations could cause a new cancer syndrome that is characterized by
mesothelioma and UM. Recently, it has been widely proven that mutation of bap
Bap1 is presumed
to be a tumour suppressor gene, is located on chromosome 3p21.1, and usually
undergoes an inactive mutation of one copy and deletion of the other copy with
the loss of one chromosome 3[13]. Dey et al[14] found that deletion of the bap1 gene in mouse
was lethal during embryogenesis, but systemic or haematopoietic-restricted
deletion in adults demonstrated features of human myelodysplastic syndrome. At
the cellular level, deficiency of BAP
The BAP1
protein is a member of the ubiquitin C‑terminal hydrolase (UCH) subfamily of
deubiquitylating enzymes[7] and serves as a
regulator in maintaining the balance of the ubiquitination cycle of histone H
In this
study, we first screened and confirmed a new BAP1 protein partner, calpastatin
(CAST), by means of protein chip, immunoprecipitations (IPs) and surface
plasmon resonance (SPR) analysis. CAST is an inhibitor of calpain, which plays
an important role in cell migration. Thus, we further explored the functional
interaction between BAP1 and CAST in cell migration and motility. We
demonstrated that CAST might play a key role in BAP1-related cell migration
regulation in UM cells.
MATERIALS AND METHODS
Cell Lines
and Cell Culture Human UM OCM
Transfection
and Lentiviral Infection For the
knockdown assay, lentiviral-based short hairpin RNA (shRNA; Obio Technology,
Shanghai, China) was applied to deplete BAP1 or CAST. Lentiviral pLKD-eGFP
shRNA vectors expressing the shRNA sequence against BAP1 (NM_004656.2, target
sequence: CGTCCGTGATTGATGATGATA), CAST (NM_001042440, target sequence:
GCTCGACCTCCGC TCAATTAA) and control (target sequence: TTCTCCGAA CGTGTCACGT)
were constructed. In the overexpression experiments, CAST (pLenti-EF
Cell
Migration Assays Wound
healing assays were carried out in these cell lines by plating 4×105 cells
in 6-well plates overnight. Before scratching with a 200 μL tip, culture media
was replaced with serum-free media. A live cell imaging system with a Leica
microscope (Leica Microsystems, DMZ6000B, Germany) was applied to capture
images (200×) every hour for 24 or 72h, and the scratch was measured using
ImageJ.
In the
transwell assay, suspended in serum-free medium, 150 μL of cells (1.0×105
cells/mL) was added to the upper chamber, and 600 μL of medium with 20% FBS was
added to the lower chamber. After 24h, the cells from the upper side of the
chamber were removed, and the chambers were soaked in crystal violet solution
to stain the cells on the lower side for 20min. Subsequently, the chambers were
washed three times with ddH2O. When the chambers were dried, photos
were taken of five random fields of view for every group using a microscope
(Leica DMI3000B, Germany). Finally, the cell numbers were counted using ImageJ.
Experiments were independently repeated three times.
Tracking the
Migration of Tumour Cells With minor
modification from a previously described protocol[19],
eGFP images of tumour cells were acquired in a live cell imaging system. After
being placed in 6-well plates, 5×104 cells were cultured overnight
at
Antibodies
and Proteins The primary
antibodies used were mouse anti-human BAP1 antibody (Santa Cruz, Texas, USA;
sc-28383), mouse anti-human calpastatin antibody (CAST, Santa Cruz; sc-20779),
rabbit anti-human vinculin antibody (Santa Cruz; sc-5573) and rabbit anti-human
calpain antibody (Santa Cruz; sc-30064). The secondary antibodies included goat
anti-mouse antibody (Santa Cruz; sc-2005) and goat anti-rabbit antibody (Santa
Cruz; sc-2004). The human full-length proteins of BAP1 (Abnova, Taipei, Taiwan,
China, H00008314-P01; Abcam,
Cambridge, UK, ab188681) and calpastatin (Abcam, ab112256) were used in
microarrays or SPR.
Human
Proteome Microarray To screen
for proteins interacting with BAP1, we used the HuProtTM microarray
(CDI Laboratories, Inc., Mayaguez, Puerto Rico), which is composed of
approximately 20 000 human full-length proteins with N-terminal glutathione
S-transferase (GST) tags. The HuProtTM microarray was performed
according to the following procedure, as previously described[20]. The human full-length BAP1 protein (Abnova) was
first concentrated with an Amicon® Ultra 3kD super filter
(Millipore), and the buffer solution was replaced with labelling buffer (Full
Moon protein-labelling kit) to remove tris. Next, biotin-labelled human BAP1
recombinant protein was prepared for testing. The microarray was blocked for
1.5h in blocking buffer [5% bovine serum albumin (BSA) in 1×phosphate buffered
saline with 0.1% Tween 20 (PBST), pH 7.2] and then incubated with BAP1 protein
sample at a final concentration of 1 µg/mL in blocking buffer overnight at
Immunoprecipitations
and Western Blots To verify
the interactions between BAP1 and selected proteins suggested by the proteome
microarray in cells under physiological conditions, IP was performed using an anti-BAP1
antibody. OCM
For Western
blots, cells were lysed and centrifuged to obtain the extracts as previously
described. For each sample, 30 μg of protein was loaded and subjected to
SDS-PAGE for 2h. Next, the samples were transferred to a PVDF membrane and
blocked in TBST (20 mmol/L Tris, pH 7.5. 150 mmol/L NaCl, 0.1% Tween 20) with
5% skim milk for 1h. The membranes were incubated with primary antibodies
overnight at
Surface
Plasmon Resonance Analysis To further
determine the real-time data on affinity and interaction kinetics between two
proteins (BAP1 and CAST), SPR analysis was performed using a ProteOnTM
XPR36 protein interaction array system (Bio-Rad, Hercules, California, USA).
According to the manufacturer’s instructions, channel surfaces of the chip
(BIO-RAD, ProteOnTM Sensor Chip, GLC 176-5011) were activated by
injection of the amine coupling reagents 1-ethyl-3-(3-dimethylaminopropyl)
carbodiimide hydrochloride (EDAC, 100 mmol/L) and N-hydroxysulfosuccinimide
(sulfo-NHS, 25 mmol/L) (components of the ProteOn amine coupling kit). The
human full-length CAST protein was immobilized onto the 1600RU channel after
being diluted to 50 ng/μL in 10 mmol/L NaAc solution (pH 4.5). To deactivate
the remaining carboxyl groups in the CAST and blank channels, 1 mol/L
ethanolamine HCl, pH 8.5 (ProteOn amine coupling kit), was then injected at a
flow rate of 30 μL/min for 300s. The channels were washed twice with running
buffer (10 mmol/L PIPES, 150 mmol/L NaCl, 0.005% Tween 20, pH
6.0). Serial dilutions of BAP1 (Abcam) samples were prepared at 160, 64, 25.6,
10.24, 4.096, and 1.638 nmol/L in PIPES solution. Samples (400 µL) of each
concentration were injected into the analyte channels orthogonal to the CAST
and blank channels at a flow rate of 50 μL/min. The binding kinetics for the
interactions of CAST and BAP1 were then rapidly and accurately obtained.
Calpain
Activity To assess
the activity of calpain in intact cells, we used a cell-permeable synthetic
fluorogenic substrate for calpain, N-Succinyl-Leu-Leu-Val-Tyr
7-amido-4-methylcoumarin (suc-LLVY-AMC, Sigma, Missouri, USA; S6510)[21]. The intact substrate exhibits little fluorescence at
445 nm upon excitation at 345 nm. However, specific proteolysis of the
substrate by calpain emits the fluorescent AMC group, leading to an increase in
its fluorescence. After infection, the cells were washed with Hank’s balanced
salt solutions (HBSS) followed by digestion and suspension at 1×105/mL
in HBSS. A 90 μL cell suspension was plated in 96-well plates and kept on ice
until the assay was performed. The substrate suc-LLVY-AMC was kept in dimethyl
sulfoxide (DMSO) (5 mg/mL) at
RESULTS
Loss of BAP1
Reduces Cell Migration To study the
effects of BAP1 loss in cell lines, we used shRNA to knockdown the expression
of BAP1. In the wound healing assays, of the three cell lines, HeLa and 92.1
cells in the shBAP1 group were less motile than control cells (Figure
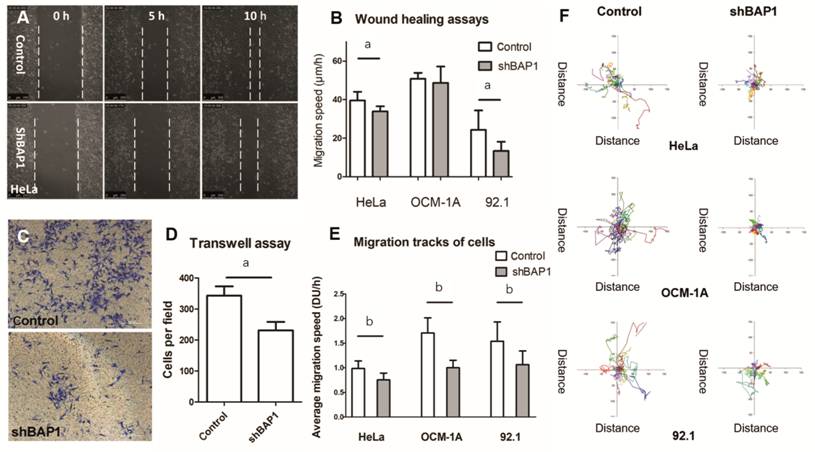
Figure 1 Loss of BAP1 reduces cell migration A, B: Representative images and quantification of
wounding healing assays of control and shBAP1 groups in three cell lines (HeLa:
human cervical cancer cell, OCM
Identification
of Proteins Interacting with BAP1 Using the Human Proteome Microarray The HuProtTM
microarray examines 47616 probes including interior labels, positive controls,
blanks and 19841 proteins, and every protein has two duplicate probes in case
of false positives. In our study, due to slight residual tris in the buffer,
the background fluorescence of the chip was slightly high. However, the
following 5 proteins were found to have an SNR>0.3 and interact with BAP1
protein: vinculin (VCL, GenBank: BC039174), CAST (NM_173060.2), phytanoyl-CoA
hydroxylase-interacting protein-like (PHYHIPL, NM_032439.1), galectin-9
(LGALS9, NM_002308.3) and von Willebrand factor A domain-containing protein
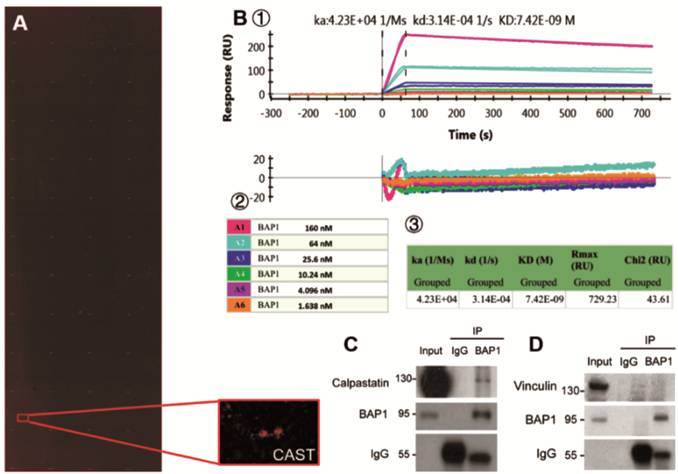
Figure 2
CAST interacts with BAP1 protein A:
Scannogram of the human proteome microarray (HuProtTM microarray)
detecting interacting proteins with BAP1 and a partially enlarged lattice
indicating the positive signal of CAST; B: Interactions between BAP1 and CAST
proteins by surface plasmon resonance analysis:① the
original graph (upper) and the weighted graph (lower) showing the binding
kinetics for the interaction between CAST and BAP1 over time; ② legend
showing dilutions of BAP1 protein; ③ table
listing the dissociation constant (KD value, 7.42×10-9) between
these two proteins; C, D: IPs in OCM
BAP1
Interacts with Calpastatin To verify
the interactions between BAP1 and VCL or CAST identified by proteome microarray
under physiological conditions, IP in OCM
Loss of BAP1
Reduces the Expression and the Function of Calpastatin Since BAP1
is a deubiquitinase, which is usually involved in regulating the protein level
of downstream targets, we tested whether BAP1 regulates the CAST protein level.
By using immunoblotting, we found that knockdown of BAP1 significantly
decreased the protein level of CAST in two UM cell lines (Figure
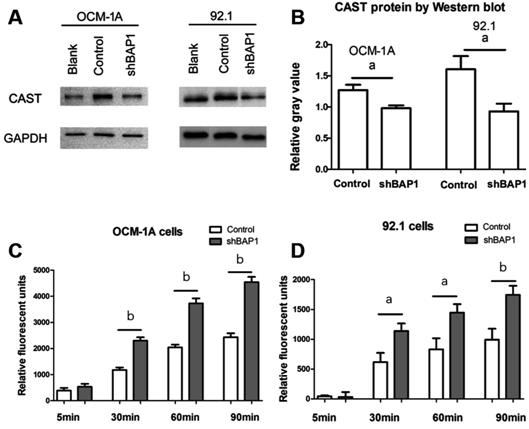
Figure 3
Loss of BAP1 decreased the expression and function of CAST A, B: CAST expression by Western blot
between control and shBAP1 groups in OCM
Calpastatin
Rescues the Cell Migration Phenotype Caused by BAP1 Loss To directly
test our hypothesis that CAST is a downstream target of BAP1 mediating its
ability to regulate cell migration, we performed a rescue experiment by
overexpressing CAST in 92.1 cells with BAP1 knockdown and then analysed the
cell migration ability by wound healing and transwell assays. In the wound
healing assay, consistent with our previous results, the migration ability of
92.1 UM cells was significantly reduced by knockdown of BAP1 (Figure
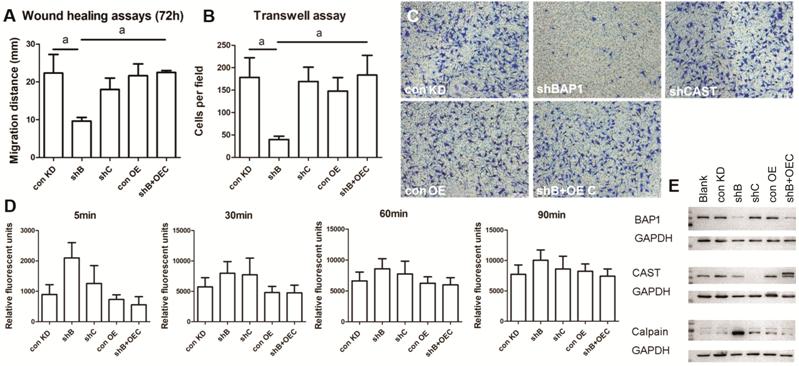
Figure 4
CAST overexpression in BAP1-deficient cells enhances cell migration A:
Wounding-healing assay of indicated groups of 92.1 cells; B, C: Transwell assay
of indicated groups of 92.1 cells with statistical results and representative
images; D: The calpain activities of indicated groups by the calpain activity
assay with Suc-LLVY-AMC; E: Protein levels of BAP1, CAST and calpain in 92.1
cells transduced with the indicated shRNAs were analysed by Western blot. con
KD: Control for knockdown; shB: shBAP1; shC: shCAST; con OE: Control for
overexpression; shB+OEC: shBAP1+overexpressing CAST. n=3. Data are
presented as the mean±SEM. Differences between groups were assessed by unpaired
t-test with/without Welch’s correction. aP<0.05.
To further
confirm the rescue effect of CAST, we analysed its activity by calpain activity.
Consistent with our previous data, knockdown of BAP1 increased calpain
activity, suggesting that CAST is dysfunctional. However, overexpression of
CAST in these cells fully rescued the calpain activity to the control level.
Since CAST also inhibits calpain at the expression level, we further analysed
the calpain expression level by Western blot. Consistent with a previous study[22-23], knockdown of CAST increased the
calpain expression level (Figure 4E). We also found that the calpain expression
level was significantly increased in the BAP1-deficient cells (Figure 4E),
which suggests a reduction in CAST function and is consistent with our
hypothesis that BAP1 is necessary for CAST function. Interestingly, however,
the calpain level upon knockdown of CAST was significantly lower than that upon
BAP1 knockdown (Figure 4E), suggesting that additional pathways contribute to
BAP1-regulated calpain expression. After overexpression of CAST in these cells,
the calpain expression level again dropped to the control level (Figure 4E),
suggesting that CAST rescued the enhanced calpain activity in the
BAP1-deficient cells.
Taken
together, our data suggest that CAST and its subsequent calpain pathway may
mediate BAP1-regulated cell migration.
DISCUSSION
The bap1
mutation has been recognized as an indicator of poor prognosis in UM[6-7,13]. However,
knockdown of BAP
In this
study, we employed a knockdown model of BAP1 to detect its possible function
and underlying mechanism in regulating cell migration. Based on the protein
chip, IP and SPR analysis, we found that BAP1 can interact with CAST in UM
cells. As an inhibitor of calpain, CAST suppresses the expression and activity
of calpain[21-23]. Calpain is
a calcium-dependent, soluble, neutral, protease that promotes cell motility
with hydrolysis of specific substrates in integrin activation, adhesion complex
turnover and membrane protrusion dynamics[23].
Calpain-mediated migration and invasion mechanisms include focal adhesion
turnover, reinforcing the expression and activity of matrix metalloproteinases
(MMP)1, MMP2 and urokinase plasminogen-type activators (uPAs), protein tyrosine
phosphatase 1B (PTP1B)-SRC-mediated invadopodia formation, cortactin-mediated
actin reorganization, as well as lamellipodia and pseudopodia stabilization at the
leading edge of the cell[24]. Under
calpain-mediated regulation of the cytoskeleton, cells can stretch and migrate
similar to amoeba. Thus, we hypothesized that CAST might participate in
cellular migration of bap1-mutated UM cells.
First, we
found the biological protein binding effects between BAP1 and CAST by means of
proteome microarray and SPR and further verified the interactions under
physiological conditions in UM cells using IP (Figure 2). Next, when we knocked
down BAP1, CAST expression decreased, and the activity of calpain increased
(Figure 3). Lastly, we overexpressed CAST in cells after BAP1 knockdown and
found that the cell migration capacity in this group was significantly enhanced
compared to the shBAP1-only group and restored fully to the control level. Our
data suggest that BAP1 mediates cell migration by downregulating the CAST
protein level and function. We also noticed that knockdown of CAST alone did
not increase the cell migration ability. These data suggest that reductions in
CAST protein level and function in BAP1-deficient cells alone may not be
sufficient to alter cell migration behaviour and additional pathways that also
change upon BAP1 loss are required to amplify such defects. However,
overexpression of CAST fully restored the cell migration defect induced by loss
of BAP1, suggesting that CAST dysregulation may serve as an initial but key
step in a sophisticated cascade upon loss of BAP1 that eventually leads to
altered cell migration. Consistent with this hypothesis, we noticed that BAP1
knockdown alone significantly increased the calpain level beyond the CAST
knockdown level, suggesting that additional factors related to BAP1 contribute
to this pathway, which needs to be further characterized.
In summary,
we confirmed that the loss of BAP1 could inhibit cellular migration capacity in
UM cell lines. Moreover, for the first time, we identified that CAST is a
strong BAP1 interacting partner. We also found that CAST plays a key role in
BAP1-related cell migration and that the overexpression of CAST fully rescued
BAP1-induced cell motility defects. Thus, our data supports a novel mechanism
underlying the cellular function of BAP1 and may shed light on the pathological
role of BAP
ACKNOWLEDGEMENTS
Foundation: Supported by
the Science and Technology Commission of Shanghai (No.14411961800).
Conflicts of
Interest: Yue H, None; Meng FX, None; Qian J, None; Xu
BB, None; Li G, None; Wu JH, None.
REFERENCES